Devyser announces study publication demonstrating dd-cfDNA detection in dual donor kidney transplant patients
Devyser Diagnostics AB announces today an article titled “Detection of donor-derived cell-free DNA...
Post-transplantation
Post-transplantation | February 7, 2023
Improving chimerism measurement in post-transplant patient monitoring
Relapse of the underlying malignant disease is a major complication after hematopoietic stem cell transplantation (HSCT). Clinicians need reliable and sensitive methods to detect relapse as early as possible and trust the results of the test used. Different methods to determine chimerism have been developed and are currently used in laboratories worldwide. Here we present the characteristics and performance of Devyser Chimerism, an NGS-based chimerism method with unmatched sensitivity.
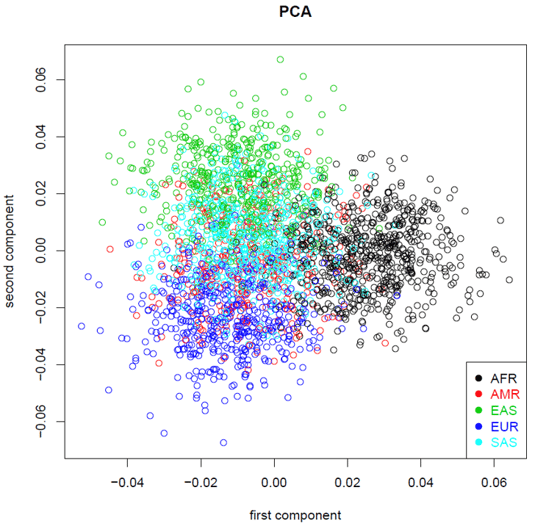
Population-independent highly informative genetic markers
The Devyser Chimerism NGS kit for the detection of mixed chimerism employs highly informative genetic markers distributed through the human genome. The markers have been selected to be population-independent (figure 1). The kit enables screening of donors and recipients prior to HSCT, as well as quantitative determination of mixed chimerism in the recipient following HSCT.
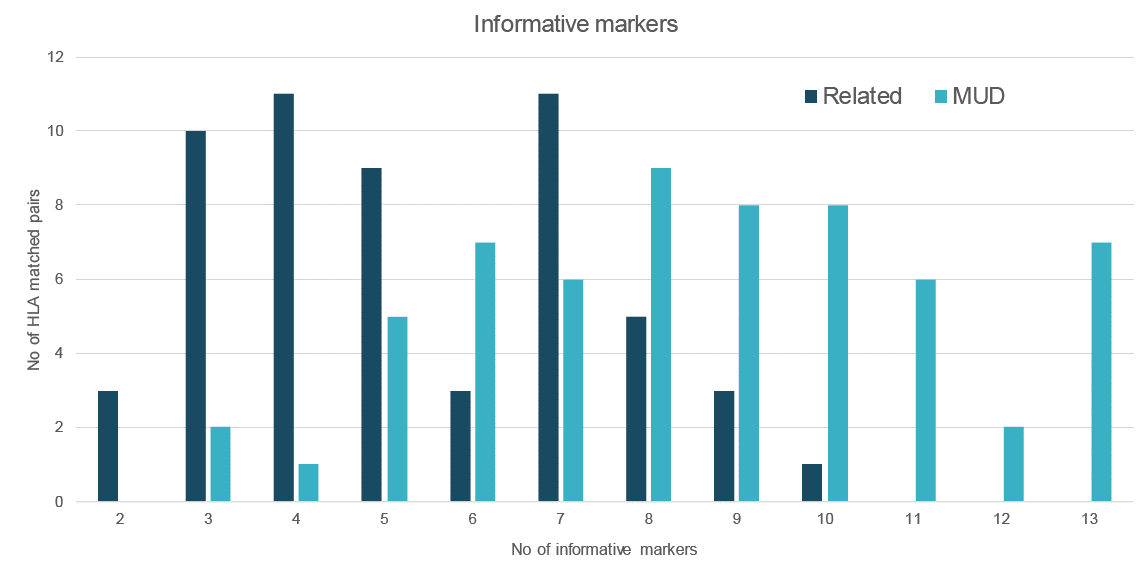
Informative markers
When testing screened patients, Devyser Chimerism exhibited at least three (average eight) and at least two (average five) informative genetic markers (indels), suitable for monitoring mixed chimerism of patients with their corresponding matched unrelated or related donor samples (Fig. 2)
A sensitive and precise assay
When comparing Devyser Chimerism to other methods, real-time PCR although very sensitive, has poor precision above 20% chimerism. In comparison, fragment analysis exhibits good precision, but with limited sensitivity (> 2,5%). In contrast, Devyser Chimerism demonstrates very good sensitivity, with a limit of detection (LOD) of 0.1% chimerism, and precision throughout the whole spectrum of patient/donor mixed chimerism (figure 3).
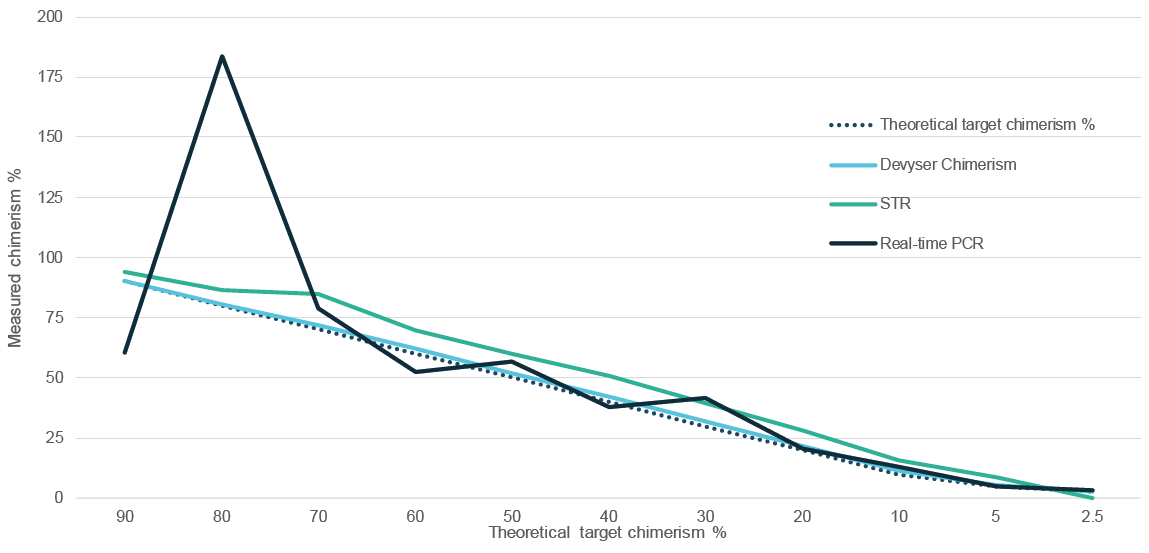
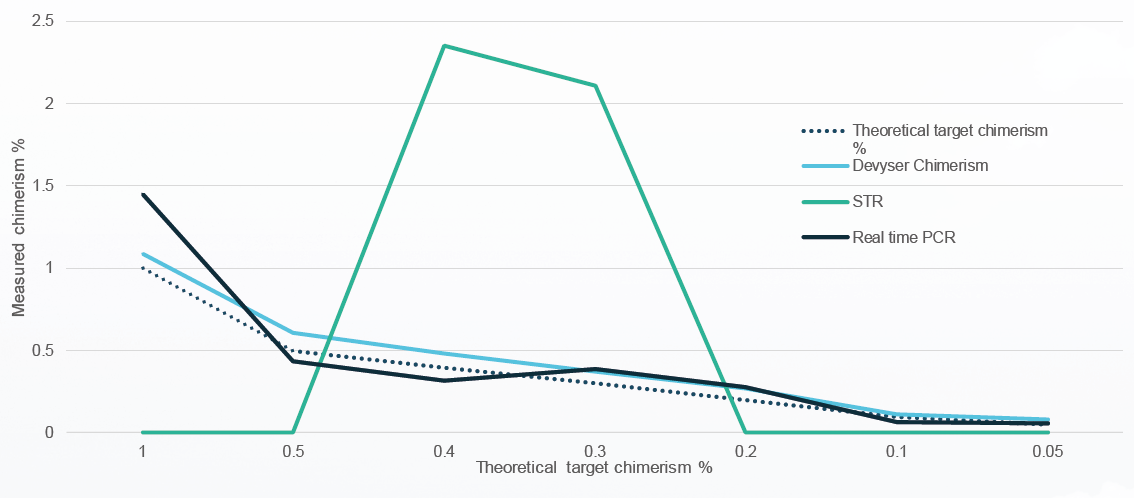 Figure 3. Devyser Chimerism compared to STR and Real-time PCR analysis (low level). STR analysis failed at 1 and 0,5%
Figure 3. Devyser Chimerism compared to STR and Real-time PCR analysis (low level). STR analysis failed at 1 and 0,5%
Conclusion
The results of our studies showed that the Devyser Chimerism assay is suitable for monitoring patient/donor chimerism after allogeneic hematopoietic stem cell transplantation (HSCT), with high sensitivity (LOD 0,1 %) and a broad dynamic range (detection range 0,1-100%) with good precision and accuracy throughout the whole spectrum of chimerism (% patient/donor)
Devyser Chimerism employs non-population dependent, highly informative genetic markers which provide stable resolution power. By utilizing the CE-IVD Devyser Chimerism kit, early rejection detection with trusted results is made possible.
Sources
.png)
Devyser Diagnostics AB announces today an article titled “Detection of donor-derived cell-free DNA...
Read More

Read More

Devyser’s novel test for detecting donor-derived cell-free DNA in blood samples from...
Read More
.jpg)
A better patient care is the final objective for Dr. Helena Devos and the team at Sint Jan’s...
Read More