Devyser launches the first IVDR compliant NGS assay for thalassemia and sickle cell disease testing
Devyser, today announced the launch of Devyser Thalassemia v2, the first Next-Generation Sequencing...
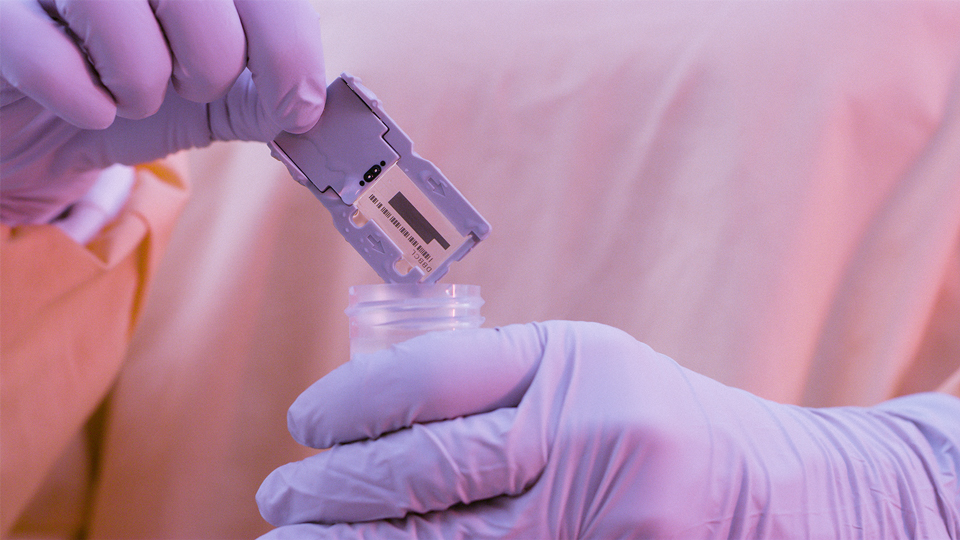
News | November 28, 2018
For many genes, choosing the most optimal transcript for a specific analysis is a complex task. This is due to two main factors. Firstly, the frequently large number of transcripts listed for a certain gene, and also because gene browsers are inconsistent in the definition of which transcripts are considered to be “default”. NCBI (RefSeq) and EMBL-EBI (Ensembl/GENCODE) are therefore working on a joint initiative to rationalise differences across gene sets, and to harmonise a subset of transcripts per gene.
The goal of the project is to choose joint “default” transcripts, so called MANE transcripts (Matched Annotations from NCBI and EBI), for which the RefSeq and Ensembl identifiers are synonyms for the same transcript feature. This dataset is to be considered a starting point for gene analysis and will provide a common framework for clinical reporting, comparative genomics, and cross-study data comparisons.
Prerequisites for inclusion as a MANE transcript
Must match GRCh38 reference sequence
One transcript per locus
100% identity between RefSeq and corresponding Ensembl transcript in CDS (coding sequence) and 5´+ 3´ UTR (untranslated region) ends
Well-supported, expressed and conserved
Representative of biology at each locus
Fairly stable over time
Preferably supported by clinical data
In this process, the selection of default transcripts is conducted automatically using two concurrent computational pipelines that compare all annotated transcripts in RefSeq or Ensembl respectively. The pipelines employ slightly different selection criteria and weighting, and were developed independently, to allow validation of the results.
The goal for the initiative is to define a “MANE-default” transcript for 50% of all genes in 2018 and 90% in 2019 and to have the MANE-transcript defined as default across all genome browsers. In addition, work is ongoing with UniProt to highlight the corresponding protein isoform as canonical. The selected MANE transcripts should be fairly stable, but updates will be considered as new data sets are uncovered and refined.
For more information about the MANE Transcripts project, including selection strategy and decision making approaches, we recommend reviewing the MANE-project blog.
.jpg)
Devyser, today announced the launch of Devyser Thalassemia v2, the first Next-Generation Sequencing...
Read More
Devyser Diagnostics AB (publ), a Swedish molecular diagnostics company, today announces that it has...
Read More

Devyser today announced that it entered a strategic agreement with Illumina, a global leader in DNA...
Read More

Devyser is proud to announce that the company has been awarded a tender by Oslo University Hospital...
Read More