Devyser launches the first IVDR compliant NGS assay for thalassemia and sickle cell disease testing
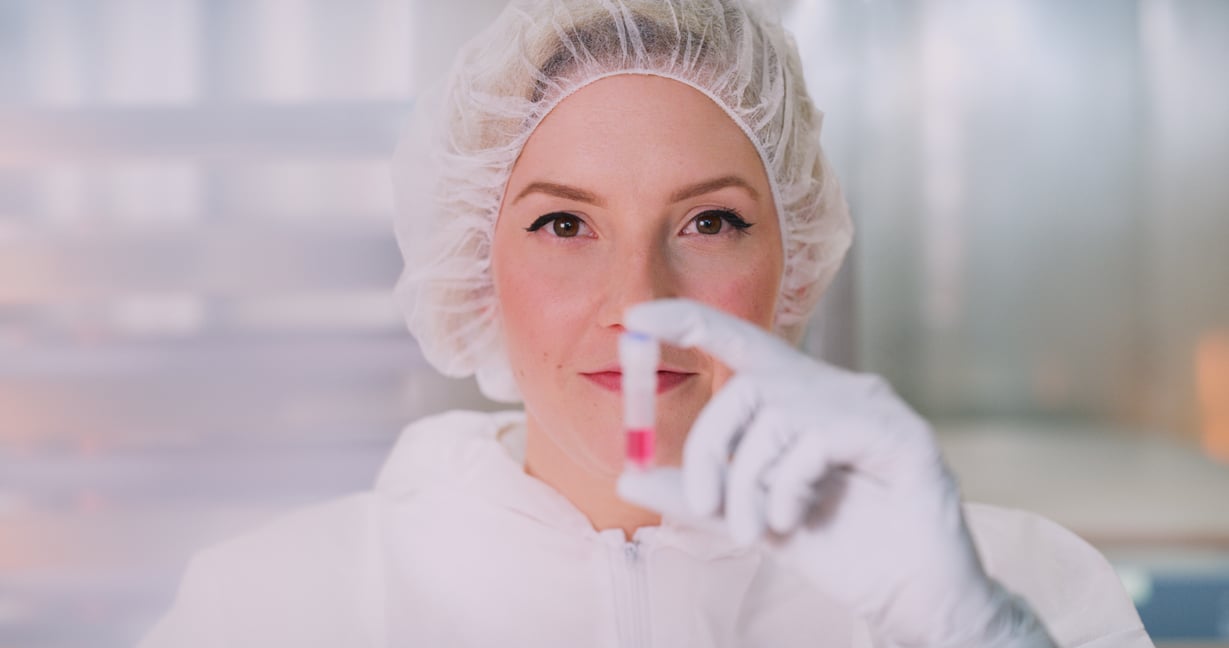
Devyser, today announced the launch of Devyser Thalassemia v2, the first Next-Generation Sequencing...

News | July 19, 2018
Two new studies attempting to define the genetic landscape of individuals who later develop Acute Myeloid Leukemia (AML) were recently published in Nature and Nature Medicine
Nature Medicine published a report (P Desai et al) regarding 212 women from the Women’s Health Initiative who were healthy at study baseline, but eventually developed AML during follow-up (median time: 9.6 years). Deep sequencing on peripheral blood DNA of these cases was performed and compared to age-matched controls that did not later develop AML. The authors discovered that mutations in IDH1, IDH2, TP53, DNMT3A, TET2 and spliceosome genes significantly increased the risk of developing AML.
Nature also published a study (S Abelson et al) in which deep sequencing of genes that are recurrently mutated in AML was performed in order to distinguish between individuals who have a high risk of developing AML and those with benign ARCH (age-related clonal haematopoiesis).
The research group analysed peripheral blood cells from 95 individuals that were obtained on average 6.3 years before AML diagnosis, together with 414 unselected age- and gender-matched individuals (control group). Pre-AML cases were distinct from controls and had more mutations per sample, higher variant allele frequencies, indicating greater clonal expansion, and showed enrichment of mutations in specific genes. This group subsequently developed an AML predictive model using health record database that identified individuals at greater risk for developing AML.
References:
Somatic mutations precede acute myeloid leukemia years before diagnosis, P Desai et al, Nature Medicine volume 24, pages1015–1023 (2018).
Prediction of acute myeloid leukaemia risk in healthy individuals, S Abelson et al, Nature, 09 July 2018.
.jpg)
Devyser, today announced the launch of Devyser Thalassemia v2, the first Next-Generation Sequencing...
Read More
Devyser Diagnostics AB (publ), a Swedish molecular diagnostics company, today announces that it has...
Read More

Devyser today announced that it entered a strategic agreement with Illumina, a global leader in DNA...
Read More

Devyser is proud to announce that the company has been awarded a tender by Oslo University Hospital...
Read More