Devyser launches the first IVDR compliant NGS assay for thalassemia and sickle cell disease testing
Devyser, today announced the launch of Devyser Thalassemia v2, the first Next-Generation Sequencing...

Oncology | September 30, 2022
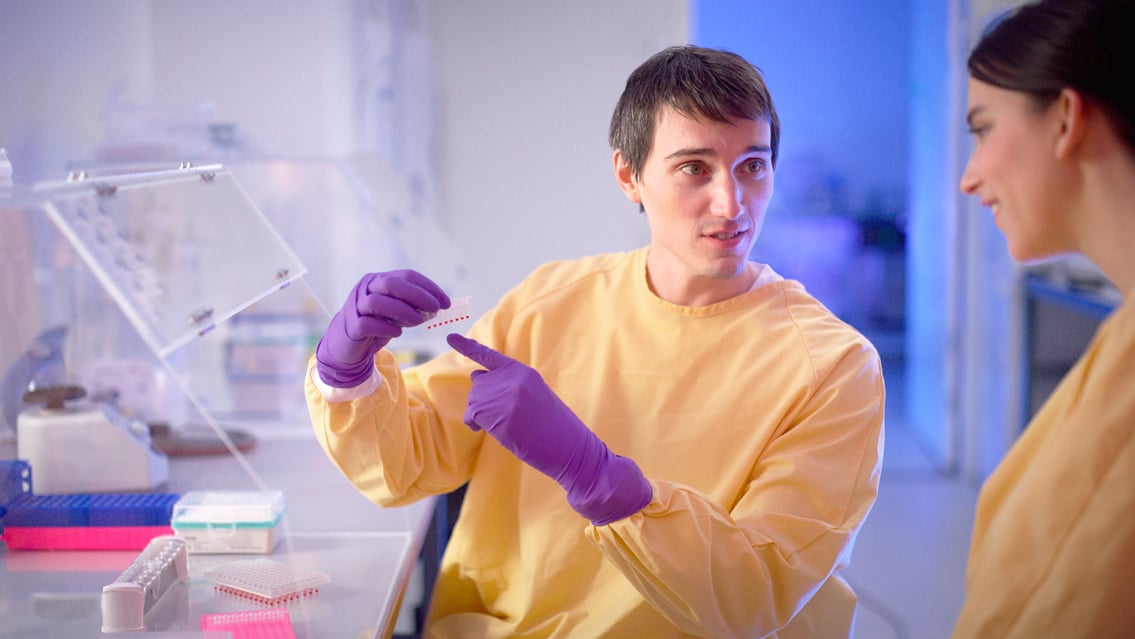
Do you know a man who has participated at least once in “Movember”? Every year, thousands of men participate in this event to raise awareness for mens prostate and testicular cancers, as well as mental health.
But what does the identification and treatment path look like for men with cancer? It turns out, much the same as for women.
It’s well known that breast cancer can also affect men (although a lesser number than women), but recent research has indicated that up to 30% of men with the most difficult-to-treat varieties of prostate cancer, also possess mutations in the BRCA1 and BRCA2 genes. An association with the ATM gene, in which mutations are also linked to increased risk of breast cancer and melanoma, was also found.
Early identification of these genes may allow for more targeted treatment. One recent study of men undergoing treatment for prostate cancer with the drug olaparib found that the response (limiting progression of their cancer) was more positive (28%) in those possessing mutations in BRCA1, BRCA2 or ATM genes, than in those without (22%). PARP-inhibiting drugs have been an approved treatment for BRCA1 and BRCA2 deficient breast and ovarian cancers since 2014, but the link between these genes and precision medicine for treating prostate cancer in the same way is still undergoing clinical evaluation.
Nonetheless, it seems clear that early identification of mutations in these genes can be just as significant for male cancer patients as female cancer patients, allowing doctors to better apply treatment plans with the greatest chance of reducing or eliminating the progress of cancers.
Devyser’s NGS oncology products provide fast, reliable, and replicable results for the identification of germline and somatic mutations.
.jpg)
Devyser, today announced the launch of Devyser Thalassemia v2, the first Next-Generation Sequencing...
Read More
Devyser Diagnostics AB (publ), a Swedish molecular diagnostics company, today announces that it has...
Read More

Devyser today announced that it entered a strategic agreement with Illumina, a global leader in DNA...
Read More

Devyser is proud to announce that the company has been awarded a tender by Oslo University Hospital...
Read More