Devyser launches the first IVDR compliant NGS assay for thalassemia and sickle cell disease testing
Devyser, today announced the launch of Devyser Thalassemia v2, the first Next-Generation Sequencing...
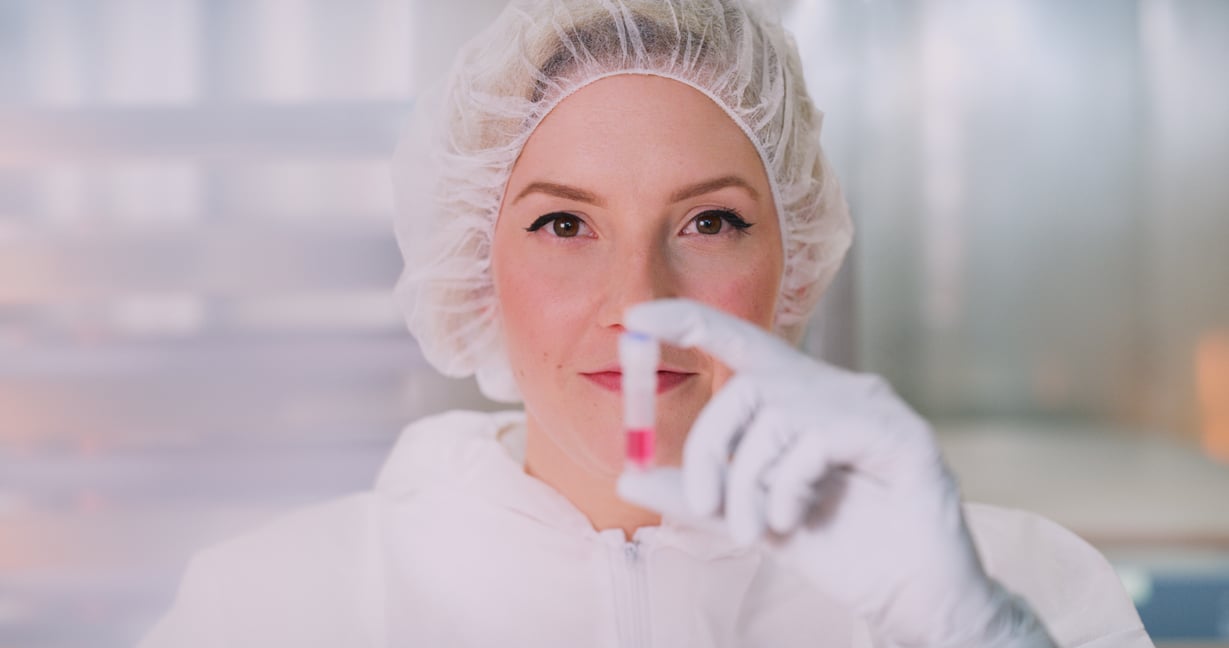

Thalassemias | October 7, 2022
Thalassemia is typically caused by sequence variants in the HBA1, HBA2 or HBB genes. The common sequence variants include a wide range of sequence variants, such as Single Nucleotide Variants (SNVs), smaller insertion and deletions (Indels) as well as large, exon spanning Copy Number Variations (CNVs). A range of different techniques such as GAP-PCR, Sanger sequencing, reverse hybridisation and MLPA is traditionally required to assess all variants.
Testing for both Alpha Thalassemia and Beta Thalassemia can be a complex process. Workflows are laboratory specific and often require the use of several different techniques to obtain a result. Relying on a patchwork of methods presents challenges such as:
.jpg)
Devyser, today announced the launch of Devyser Thalassemia v2, the first Next-Generation Sequencing...
Read More
Devyser Diagnostics AB (publ), a Swedish molecular diagnostics company, today announces that it has...
Read More

Devyser today announced that it entered a strategic agreement with Illumina, a global leader in DNA...
Read More

Devyser is proud to announce that the company has been awarded a tender by Oslo University Hospital...
Read More